Cleveland Clinic and IBM Debut New Quantum Workflow for Simulating Proteins
March 23, 2026 -- For the first time, researchers have used quantum computing to simulate the electronic structure of a protein—a demonstration made possible thanks to novel quantum-centric supercomputing (QCSC) research.
A joint Cleveland Clinic-IBM research team modeled the 303-atom miniprotein Trp-cage using a quantum-centric supercomputing workflow and an IBM Quantum Heron r2. The work is a result of progress designing and applying QCSC algorithms and workflows that combine quantum and high-performance classical computing to powerful effect.
Accurate electronic structure calculations on classical computers become increasingly challenging as system size increases. Classical methods alone can efficiently model certain aspects of protein behavior, but high-accuracy quantum-mechanical treatments of entire proteins remain impractical.
If quantum computers eventually enable accurate modeling of large, biologically relevant molecules, this will significantly impact chemical, materials science, and medical research. This work represents a step in that direction.
“I’m sort of pinching myself that we were able to do it,” said Dr. Kenneth Merz, PhD, who leads the Merz lab at Cleveland Clinic.Why modeling large molecules is excitingTrp-cage is useful for benchmarking computational chemistry methods. It's relatively compact for a protein, but it has features that are common to much larger molecules in biochemistry, such as a water-repelling or “hydrophobic” core and hydrogen bonding between its constituent parts, allowing it to take on more complex structures. The researchers modeled both its unfolded and folded (i.e., stretched and contracted) states.
Why modeling large molecules is exciting
Trp-cage is useful for benchmarking computational chemistry methods. It's relatively compact for a protein, but it has features that are common to much larger molecules in biochemistry, such as a water-repelling or “hydrophobic” core and hydrogen bonding between its constituent parts, allowing it to take on more complex structures. The researchers modeled both its unfolded and folded (i.e., stretched and contracted) states.
“Proving that this approach works for the Trp-cage is a Trp-cage is a step to larger molecules,” said Mario Motta, co-author of the paper.
The team was surprised at what they’d already achieved. “At first the plan was to simulate just a couple of amino acids,” Motta said. But as they tested their workflow, they found they were able to scale all the way up to Trp-cage and get meaningful results.
As these methods mature and scale, Merz said, he hopes they could support computational workflows for pharmaceutical research and related fields. He envisions a world where scientists use QCSC workflows to build databases of simulated molecular behavior. Then, when scientists need a new molecule for a particular purpose, they could use machine learning algorithms trained on those databases, asking for molecules that might behave in in the ways they need. From there, they could synthesize those molecules to test in real life.
A new workflow for simulating large molecules
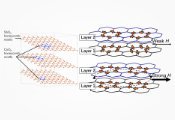
The workflow, described in a recent preprint on arXiv, relies on a technique called wave function-based embedding (EWF) to fragment Trp-cage into computationally tractable pieces called “clusters.” In EWF, there are as many clusters as there are atoms in the molecule, but each cluster is more complex than a single atom: it encompasses a local region surrounding the atom and entangled with it.
In any given protein, some pieces or clusters are going to be much more complex than others. One atom may be all the way out on the edge of the protein, at the end of a covalent bond, and only entangled to one or two neighboring atoms. In those cases, researchers can find that cluster’s electronic structure efficiently using classical computational methods. Another might be closer to the molecular core, enmeshed in a more complex web of intermolecular interactions. These larger clusters are good problems for quantum computers to solve.
Stitched back together, the results of individual cluster calculations lead to a complete solution for the electronic structure of the molecule, which describes where its electrons are and how they interact—important information that determines how the molecule behaves. This approach, quantum computers load-sharing with classical computers in hybrid workflows, is an early look at quantum-centric supercomputing in action.
A new generation of quantum-centric algorithms for HPC
Merz has watched the development of quantum computing over a period of several years. Until a few years ago, it was clear that quantum computers could offer new approaches to solving hard chemistry problems, but what those approaches would look like remained an open question.
Merz said there was something of a eureka moment when he saw a group of IBM scientists present an algorithm called sample-based quantum diagonalization (SQD).
SQD belongs to an emerging set of algorithms built for quantum-centric supercomputing, where classical and quantum resources work together to solve problems, using the strengths of both paradigms. It addresses one of the fundamental challenges of electronic structure calculations: the number of possible configurations of a molecule’s electrons grows combinatorially with the molecule’s size.
In SQD, the quantum computer samples this vast space, identifying key configurations for the classical computer to focus on. The classical computer uses the resulting information to find a solution.
After learning about SQD, Merz said, “We sort of dropped everything. I met with a few people in my group over the weekend, and we decided to just go all in on SQD.”
They began putting the algorithm through its paces, testing it on a string of smaller molecules, beginning a chain of experiments that led to this Trp-cage simulation. The results so far have been extremely promising: already in this paper, the workflow performs competitively with classical approaches, approaching the accuracy of the most computationally demanding among them.
In principle, the scientists said, the combined EWF-SQD workflow can scale far beyond Trp-cage. As molecules get larger, the task of breaking them up, calculating their most complex clusters, and stitching them back together gets more complex. But solving for the electronic structure of complex clusters is a compelling problem for quantum computers. Already, the researchers are exploring what the next step looks like, eyeing even larger molecules as their targets.
As QCSC advances, it’s important that quantum and HPC researchers work together. This work was made possible by access to HPC resources at Michigan State University and Cleveland Clinic. Other recent collaborations between IBM and HPC leaders like RIKEN have also yielded exciting results.